 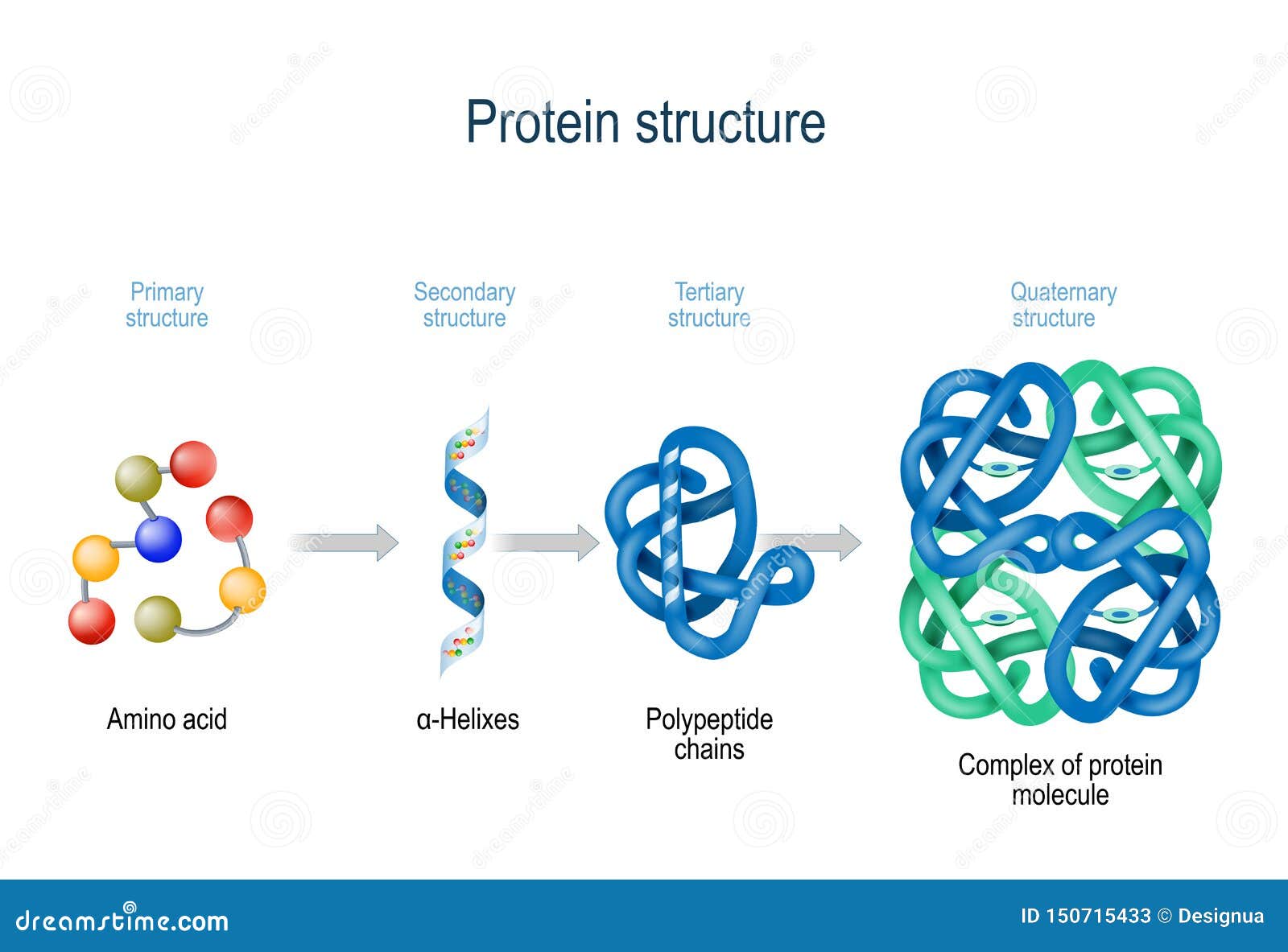
The major class of secondary metabolites from C. This report can contribute to the understanding of the structure, physicochemical properties, structural motifs, and protein-protein interaction of CURS1, CUR2, and CURS3. /protein-structure-373563_final11-5c81967f46e0fb00012c667d.png)
Prediction of the 3D model of a protein by in silico analysis is a highly challenging aspect to confirm the NMR or X-ray crystallographic data. In the predicted secondary structure of CURS proteins, alpha helices and random coils of CURS1, CUR2, and CURS3 were 42.72, 41.38, and 44.74% and 24.87, 31.03, and 17.89, respectively. Molecular weight of CURS2 and CURS3 proteins are 20266.13 Da and 20629.52 Da, theoretical pI as 5.28 and 4.96, and an aliphatic index of 89.30 and 86.37, respectively. The molecular weight of CURS1 is 21093.19 Da, theoretical pI as 4.93, and an aliphatic index of 99.19. The reliability of the modeled structure was confirmed using PROCHECK and QMEAN programs. The predicted secondary structure of curcumin synthase provided an α-helix and random coil as the major components. The primary, secondary, and tertiary structure of the CURS proteins were modeled and characterized using multiple bioinformatics tools such as ExPasy ProtParam tools, self-optimized prediction method with alignment (SOPMA), PSIPRED, and SWISS-MODEL. The present study detected the physicochemical properties and structural characteristics including the secondary and 3D structure of CURS proteins. longa is controlled by CURS1, CURS2, and CURS3 genes. Pharmaceutically important curcuminoid synthesis in C. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
